Quickstart#
This guide shows the three ways to define a sampling grid in healpix-plot, from the simplest dict shorthand to a pixel-aligned affine grid.
Setup#
import numpy as np
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
import healpix_plot
1 — Dict shorthand#
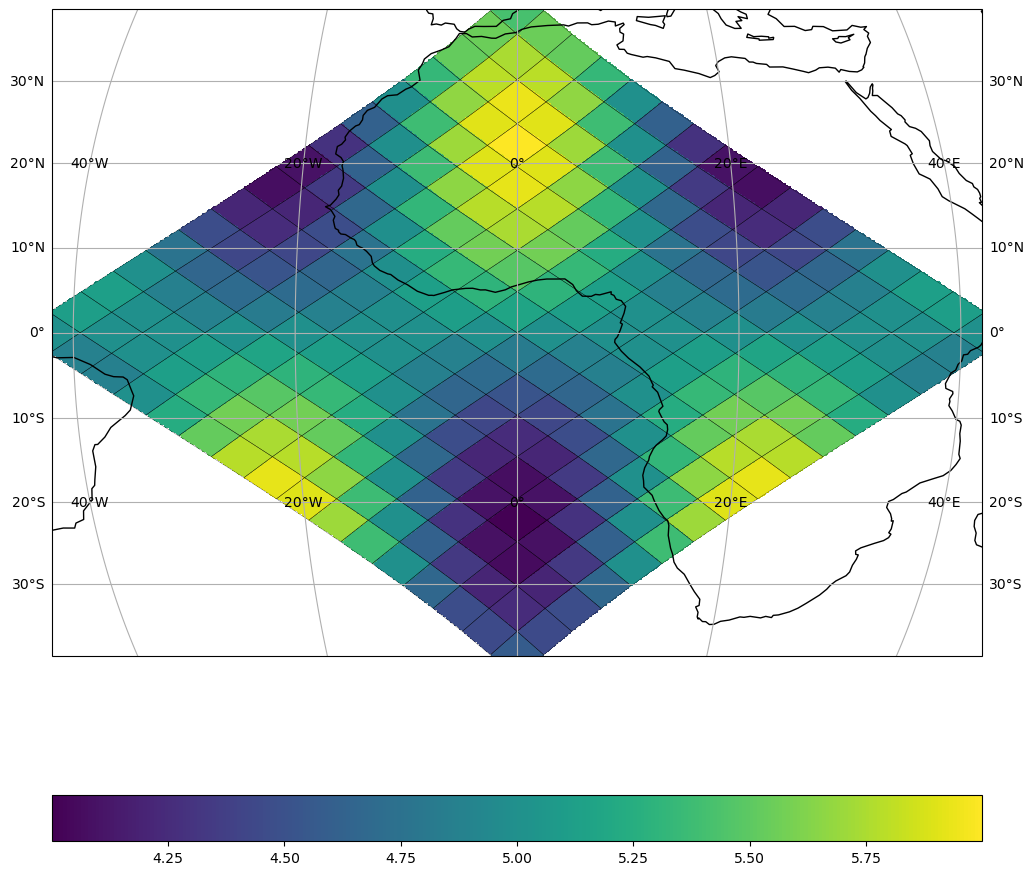
The simplest way: pass {"shape": N} and the spatial extent is inferred automatically from the cell IDs.
In real workflows your cell_ids and data come from a file (NetCDF, Zarr, …). Here we use a synthetic wave pattern.
healpix_grid = healpix_plot.HealpixGrid(
level=3, indexing_scheme="nested", ellipsoid="WGS84"
)
cell_ids = np.arange(
4 * 4**healpix_grid.level, 5 * 4**healpix_grid.level, dtype="uint64"
)
lon, lat = healpix_grid.operations.healpix_to_lonlat(
cell_ids, **healpix_grid.as_keyword_params()
)
data = np.cos(8 * np.deg2rad(lon)) * np.sin(4 * np.deg2rad(lat))
print(f"level: {healpix_grid.level}, n_cells: {cell_ids.size}")
print(f"data range: [{data.min():.2f}, {data.max():.2f}]")
level: 3, n_cells: 64
data range: [-0.99, 0.99]
fig, ax = plt.subplots(
1, 1, subplot_kw={"projection": ccrs.Mollweide()}, figsize=(12, 12)
)
mappable = healpix_plot.plot(
cell_ids, data, healpix_grid=healpix_grid, sampling_grid={"shape": 512}, ax=ax
)
fig.colorbar(mappable, orientation="horizontal")
ax = mappable.figure.axes[0]
ax.coastlines()
ax.gridlines(draw_labels="x")
ax.gridlines(draw_labels="y")
plt.show()
/home/runner/work/healpix-plot/healpix-plot/.pixi/envs/docs/lib/python3.13/site-packages/cartopy/io/__init__.py:242: DownloadWarning: Downloading: https://naturalearth.s3.amazonaws.com/110m_physical/ne_110m_coastline.zip
warnings.warn(f'Downloading: {url}', DownloadWarning)
2 — ParametrizedSamplingGrid.from_bbox — fixed region#
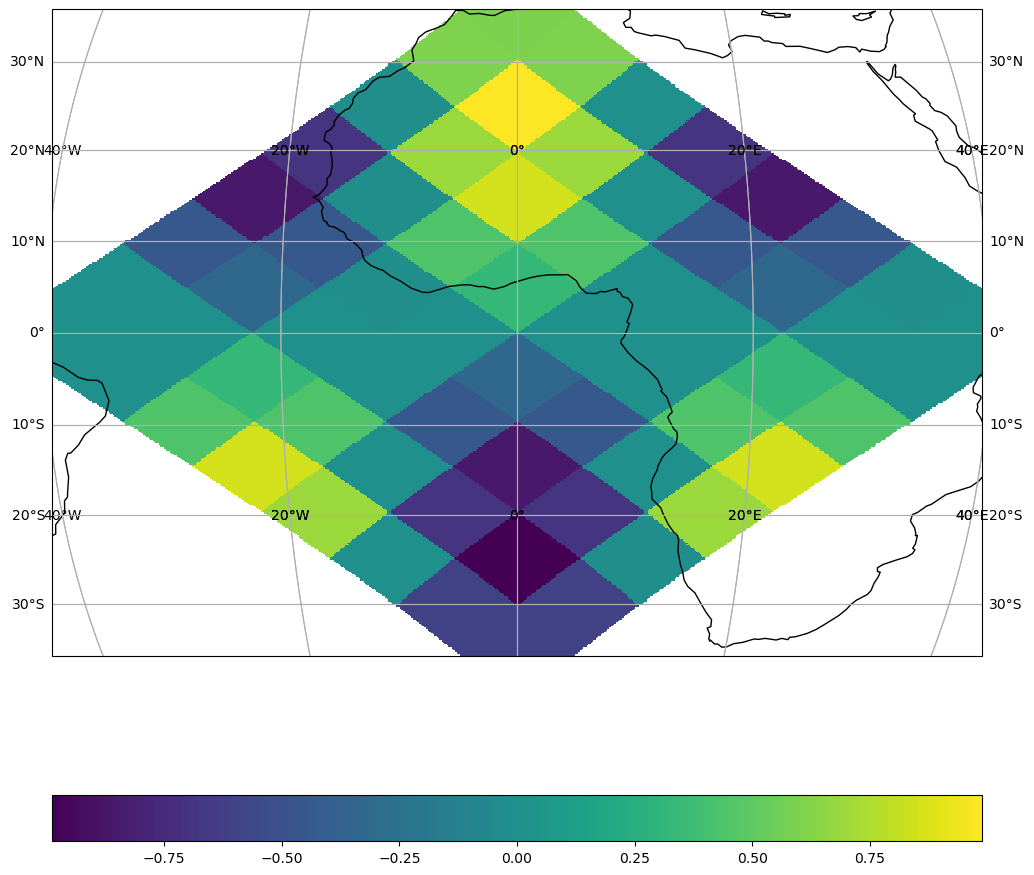
Use from_bbox when you want to pin the output to a fixed geographic extent, regardless of the data coverage. Useful for regional maps, comparisons, or animations.
Here we zoom in on the western Mediterranean.
from healpix_plot import SamplingGrid
from healpix_plot.sampling_grid import ParametrizedSamplingGrid
# Use WGS84 — the standard geodetic ellipsoid for Earth observation
healpix_grid = healpix_plot.HealpixGrid(
level=6, indexing_scheme="nested", ellipsoid="WGS84"
)
grid = ParametrizedSamplingGrid.from_bbox(
bbox=(-20, 35.0, 20, 40.0), # (xmin, ymin, xmax, ymax) in degrees
shape=512,
)
cell_ids = np.arange(
4 * 4**healpix_grid.level, 5 * 4**healpix_grid.level, dtype="uint64"
)
lon, lat = healpix_grid.operations.healpix_to_lonlat(
cell_ids, **healpix_grid.as_keyword_params()
)
data = np.cos(8 * np.deg2rad(lon)) * np.sin(4 * np.deg2rad(lat))
fig, axes = plt.subplots(
1, 1, subplot_kw={"projection": ccrs.PlateCarree()}, figsize=(12, 12)
)
mappable = healpix_plot.plot(
cell_ids, data, healpix_grid=healpix_grid, sampling_grid=grid, ax=axes
)
fig.colorbar(mappable, orientation="horizontal")
ax = mappable.figure.axes[0]
ax.coastlines()
ax.gridlines(draw_labels="x")
ax.gridlines(draw_labels="y")
<cartopy.mpl.gridliner.Gridliner at 0x7f8494b6f890>
/home/runner/work/healpix-plot/healpix-plot/.pixi/envs/docs/lib/python3.13/site-packages/cartopy/io/__init__.py:242: DownloadWarning: Downloading: https://naturalearth.s3.amazonaws.com/10m_physical/ne_10m_coastline.zip
warnings.warn(f'Downloading: {url}', DownloadWarning)
3 — AffineSamplingGrid.from_transform — pixel-aligned grid#
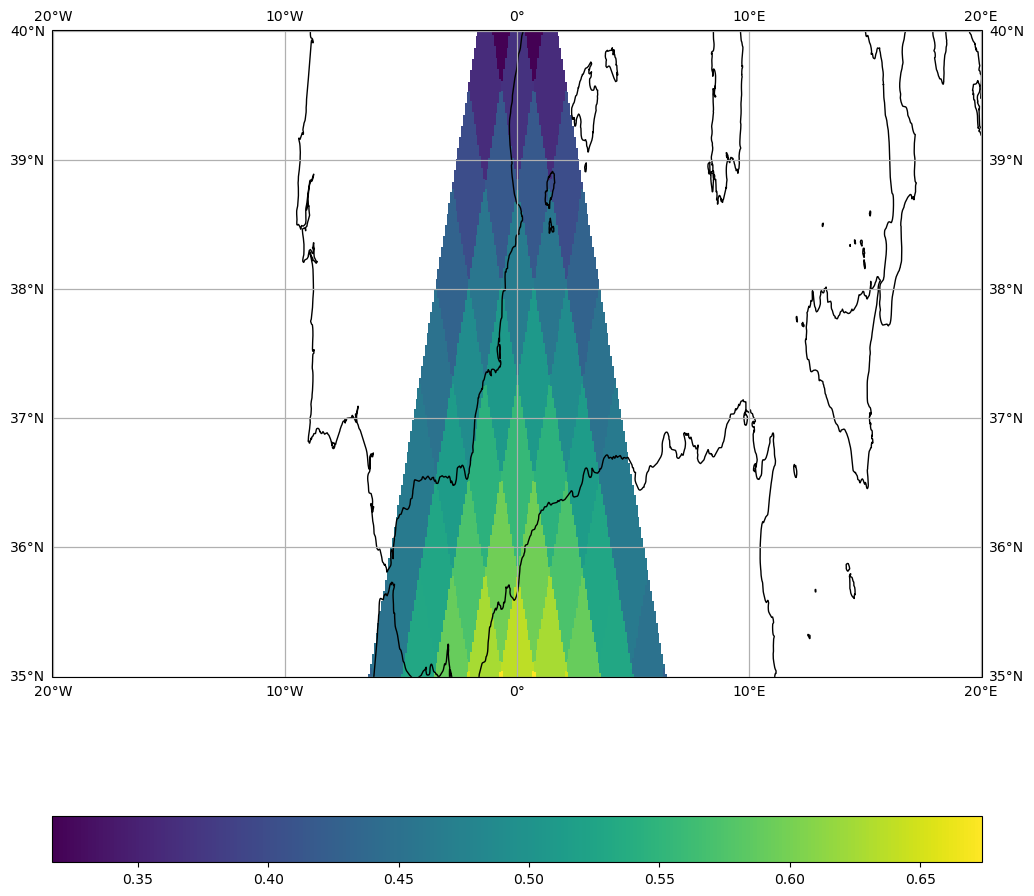
Use AffineSamplingGrid when the output pixels must align exactly with a reference raster (e.g. a GeoTIFF). The affine transform maps pixel indices to geographic coordinates: Affine(pixel_width, 0, lon_min, 0, -pixel_height, lat_max).
Here we define a 0.01°/pixel grid (~1 km) over Corsica.
from affine import Affine
from healpix_plot.sampling_grid import AffineSamplingGrid
# Use WGS84 — the standard geodetic ellipsoid for Earth observation
healpix_grid = healpix_plot.HealpixGrid(
level=6, indexing_scheme="nested", ellipsoid="WGS84"
)
cell_ids = np.arange(
4 * 4**healpix_grid.level, 5 * 4**healpix_grid.level, dtype="uint64"
)
lon, lat = healpix_grid.operations.healpix_to_lonlat(
cell_ids, **healpix_grid.as_keyword_params()
)
data = np.cos(8 * np.deg2rad(lon)) * np.sin(4 * np.deg2rad(lat))
transform = Affine(0.01, 0, -1, 0, -0.01, 40) # 0.01° resolution
grid = AffineSamplingGrid.from_transform(transform, shape=(512, 512))
fig, axes = plt.subplots(
1, 1, subplot_kw={"projection": ccrs.Mollweide()}, figsize=(12, 12)
)
mappable = healpix_plot.plot(
cell_ids, data, healpix_grid=healpix_grid, sampling_grid=grid, ax=axes
)
fig.colorbar(mappable, orientation="horizontal")
ax = mappable.figure.axes[0]
ax.coastlines()
ax.gridlines(draw_labels="x")
ax.gridlines(draw_labels="y")
<cartopy.mpl.gridliner.Gridliner at 0x7f8494e27250>
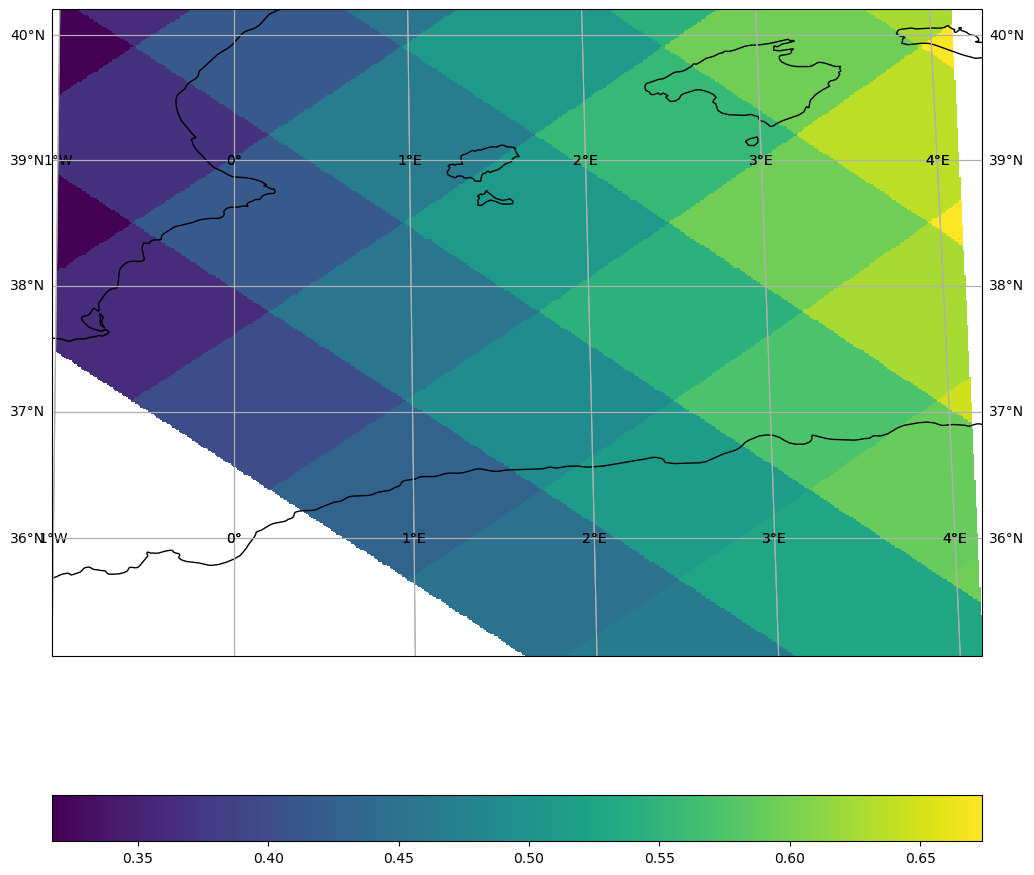
Recipe: overlay true HEALPix boundaries#
For sanity checks or presentations you can combine fast raster plots from healpix-plot with boundary polygons from xdggs: